Exploring groundwater data from Lizard
In this notebook, you will experiment how to use the hydropandas package to access, visualize and explore meta data from Lizard, a cloud datawarehouse that stores all groundwater observations from Vitens. Vitens is the largest drinking water company in the Netherlands, and it has more than 10.000 groundwater wells and more than 50.000 timeseries in its datawarehouse. The data spans from the 1930’s to the present, and it is constantly updated with new observations. Vitens also validates the
data using ArtDiver and provides quality flags and comments for each observation. The data is open to the public and you can find more information at https://vitens.lizard.net.
Feel free to customize and expand upon this introduction as needed. Happy coding! 🚀🌍📊
Notebook contents
Find groundwater wells on a map
Analyse groundwater observations
Build a Pastas model
[1]:
import logging
from IPython.display import HTML
import pastas as ps
import pandas as pd
import hydropandas as hpd
[2]:
hpd.util.get_color_logger("INFO")
[2]:
<RootLogger root (INFO)>
Get observations from extent
Use ObsCollection to find monitoring wells by specifying a geographical extent in Rijksdriehoeks coordinates.
[3]:
my_extent = (137000, 138000, 458000, 459000)
oc = hpd.read_lizard(extent=my_extent)
Number of monitoring wells: 1
Number of pages: 1
Page: 100%|██████████| 1/1 [00:00<00:00, 2.06it/s]
monitoring well: 100%|██████████| 1/1 [00:08<00:00, 8.59s/it]
Visualize all groundwater wells inside the extent on a map (visualize the ObsCollection). The markers are clickable to show a preview of the availables observations.
[4]:
oc.plots.interactive_map(color="red", zoom_start=15, tiles="Esri.WorldImagery")
[4]:
Print all the retrieved groundwater wells and tubes, and make a plot of the observations.
[5]:
oc
[5]:
| x | y | filename | source | unit | monitoring_well | tube_nr | screen_top | screen_bottom | ground_level | tube_top | metadata_available | obs | lat | lon | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| name | |||||||||||||||
| UPWP016001 | 137401.64297 | 458893.683528 | lizard | m NAP | B31H0580 | 1 | -22.43 | -24.43 | 1.58 | 2.198 | True | GroundwaterObs UPWP016001 -----metadata------ ... | 52.117985 | 5.13026 | |
| UPWP016003 | 137401.64297 | 458893.683528 | lizard | m NAP | B31H0580 | 3 | -65.43 | -67.43 | 1.58 | 2.141 | True | GroundwaterObs UPWP016003 -----metadata------ ... | 52.117985 | 5.13026 | |
| UPWP016002 | 137401.64297 | 458893.683528 | lizard | m NAP | B31H0580 | 2 | -53.93 | -55.93 | 1.58 | 2.178 | True | GroundwaterObs UPWP016002 -----metadata------ ... | 52.117985 | 5.13026 |
[6]:
oc.plots.section_plot(plot_obs=True)
INFO:hydropandas.extensions.plots:created sectionplot -> UPWP016001
INFO:hydropandas.extensions.plots:created sectionplot -> UPWP016003
INFO:hydropandas.extensions.plots:created sectionplot -> UPWP016002
[6]:
(<Figure size 1500x500 with 2 Axes>,
[<Axes: ylabel='m NAP'>, <Axes: ylabel='m NAP'>])
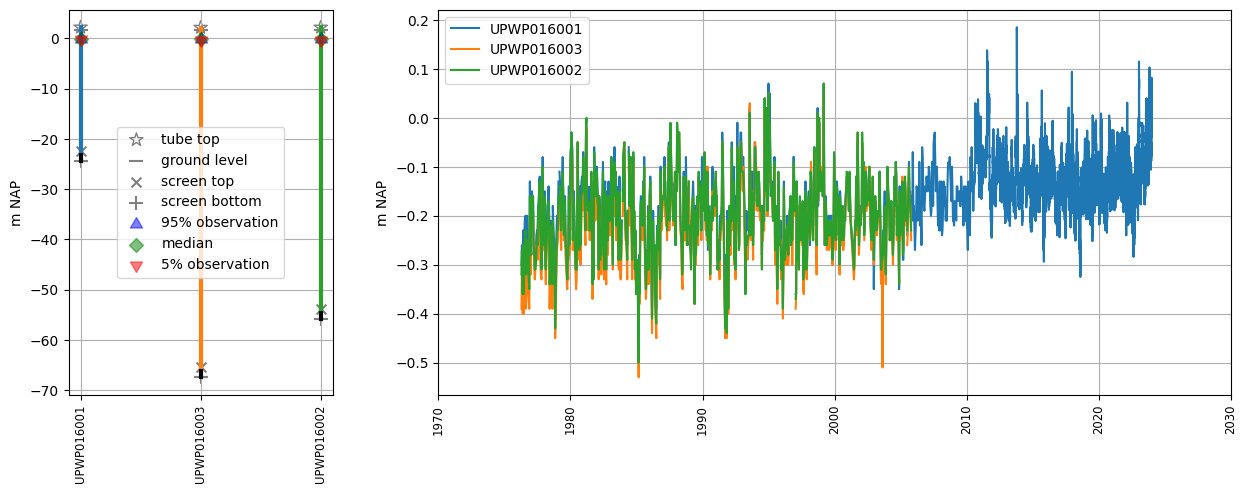
Groundwater observations
Now lets download the groundwater level observation using the from_lizard function of a GroundwaterObs object. The code below reads the groundwater level timeseries for the well UPWP016 from Lizard and makes a plot.
[7]:
gw_lizard = hpd.GroundwaterObs.from_lizard(
"UPWP016", tmin="1900-01-01", tmax="2030-01-01"
)
print(gw_lizard)
ax = gw_lizard["value"].plot(
figsize=(12, 5),
marker=".",
grid=True,
label=gw_lizard.name,
legend=True,
xlabel="Date",
ylabel="m NAP",
title="Groundwater observations for " + gw_lizard.name,
)
GroundwaterObs UPWP016001
-----metadata------
name : UPWP016001
x : 137401.64297031244
y : 458893.6835282785
filename :
source : lizard
unit : m NAP
monitoring_well : B31H0580
tube_nr : 1
screen_top : -22.43
screen_bottom : -24.43
ground_level : 1.58
tube_top : 2.198
metadata_available : True
-----time series------
value flag comment
peil_datum_tijd
1976-04-15 12:00:00 -0.300 betrouwbaar
1976-04-28 12:00:00 -0.260 betrouwbaar
1976-05-18 12:00:00 -0.330 betrouwbaar
1976-06-02 12:00:00 -0.230 betrouwbaar
1976-06-17 12:00:00 -0.300 betrouwbaar
... ... ... ...
2024-01-03 21:00:00 0.059 betrouwbaar
2024-01-04 00:00:00 0.059 betrouwbaar
2024-01-04 03:00:00 0.062 betrouwbaar
2024-01-04 06:00:00 0.066 betrouwbaar
2024-01-04 09:00:00 0.078 betrouwbaar
[29259 rows x 3 columns]
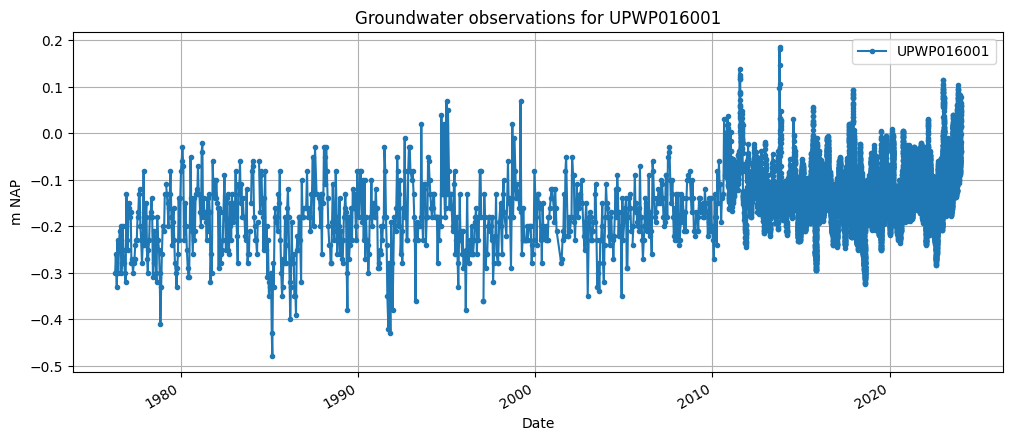
The groundwater observations contain a validation flag per timestamp. These can ‘betrouwbaar’ (reliable), ‘onbetrouwbaar’ (unreliable) en ‘onbeslist’ (unvalidated). Below flags of the timeseries are shown as a percentage, and the unreliable timestamps are printed.
[8]:
print(gw_lizard["flag"].value_counts(normalize=True) * 100)
gw_lizard[gw_lizard["flag"] == "onbetrouwbaar"]
UPWP016001
betrouwbaar 99.979493
onbetrouwbaar 0.017089
onbeslist 0.003418
Name: proportion, dtype: float64
[8]:
hydropandas.GroundwaterObs
| UPWP016001 | |
|---|---|
| x | 137401.64297 |
| y | 458893.683528 |
| filename | |
| source | lizard |
| unit | m NAP |
| monitoring_well | B31H0580 |
| tube_nr | 1 |
| screen_top | -22.43 |
| screen_bottom | -24.43 |
| ground_level | 1.58 |
| tube_top | 2.198 |
| metadata_available | True |
| value | flag | comment | |
|---|---|---|---|
| peil_datum_tijd | |||
| 1979-02-16 12:00:00 | NaN | onbetrouwbaar | Geen waarneming (-90) |
| 1983-11-29 12:00:00 | NaN | onbetrouwbaar | Geen waarneming (-90) |
| 1984-05-14 12:00:00 | NaN | onbetrouwbaar | Geen waarneming (-90) |
| 1996-12-28 12:00:00 | NaN | onbetrouwbaar | Geen waarneming (-90) |
| 2005-06-14 12:00:00 | NaN | onbetrouwbaar | Geen waarneming (-90) |
Pastas model
Lets make a Pastas model for this groundwater well (starting from 2015) and use the nearest KNMI station for meteorological data
[9]:
# Get the precipitation and evaporation data from the KNMI
precipitation = hpd.PrecipitationObs.from_knmi(
xy=(gw_lizard.x, gw_lizard.y),
start=gw_lizard.index[0],
end=gw_lizard.index[-1],
fill_missing_obs=True,
)
evaporation = hpd.EvaporationObs.from_knmi(
xy=(gw_lizard.x, gw_lizard.y),
meteo_var="EV24",
start=gw_lizard.index[0],
end=gw_lizard.index[-1],
fill_missing_obs=True,
)
# Create a Pastas Model
ml = ps.Model(gw_lizard["value"], name=gw_lizard.name)
# Add the recharge data as explanatory variable
ts1 = ps.RechargeModel(
precipitation["RH"].resample("D").first(),
evaporation["EV24"].resample("D").first(),
ps.Gamma(),
name="rainevap",
settings=("prec", "evap"),
)
# Add the stressmodel to the model and solve for period after 2015
ml.add_stressmodel(ts1)
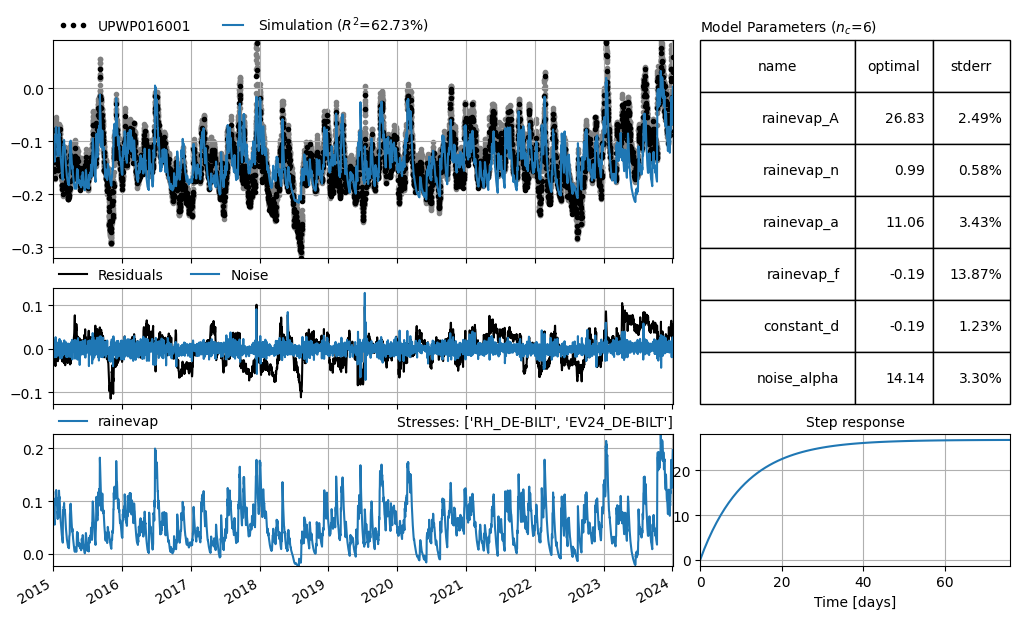
ml.solve(tmin="2015")
ml.plots.results(figsize=(10, 6))
INFO:hydropandas.io.knmi:get KNMI data from station nearest to coordinates (137401.64297031244, 458893.6835282785) and meteovariable RH
WARNING:hydropandas.io.knmi:changing end_date to 2024-01-04
INFO:hydropandas.io.knmi:get KNMI data from station nearest to coordinates (137401.64297031244, 458893.6835282785) and meteovariable EV24
WARNING:hydropandas.io.knmi:changing end_date to 2024-01-04
WARNING: The Time Series 'UPWP016001' has nan-values. Pastas will use the fill_nan settings to fill up the nan-values.
WARNING:pastas.timeseries:The Time Series 'UPWP016001' has nan-values. Pastas will use the fill_nan settings to fill up the nan-values.
INFO: Time Series 'UPWP016001': 11 nan-value(s) was/were found and filled with: drop.
INFO:pastas.timeseries:Time Series 'UPWP016001': 11 nan-value(s) was/were found and filled with: drop.
INFO: Time Series 'RH_DE-BILT' was extended in the future to 2024-01-04 00:00:00 with the mean value (0.0024) of the time series.
INFO:pastas.timeseries:Time Series 'RH_DE-BILT' was extended in the future to 2024-01-04 00:00:00 with the mean value (0.0024) of the time series.
INFO: Time Series 'EV24_DE-BILT' was extended in the future to 2024-01-04 00:00:00 with the mean value (0.0017) of the time series.
INFO:pastas.timeseries:Time Series 'EV24_DE-BILT' was extended in the future to 2024-01-04 00:00:00 with the mean value (0.0017) of the time series.
Fit report UPWP016001 Fit Statistics
=================================================
nfev 33 EVP 62.73
nobs 3291 R2 0.63
noise True RMSE 0.03
tmin 2015-01-01 00:00:00 AIC -29158.55
tmax 2024-01-04 09:00:00 BIC -29121.96
freq D Obj 0.23
warmup 3650 days 00:00:00 ___
solver LeastSquares Interp. No
Parameters (6 optimized)
=================================================
optimal stderr initial vary
rainevap_A 26.825256 ±2.49% 198.612463 True
rainevap_n 0.986379 ±0.58% 1.000000 True
rainevap_a 11.059931 ±3.43% 10.000000 True
rainevap_f -0.192514 ±13.87% -1.000000 True
constant_d -0.193830 ±1.23% -0.134993 True
noise_alpha 14.137844 ±3.30% 1.000000 True
[9]:
[<Axes: xlabel='peil_datum_tijd'>,
<Axes: xlabel='peil_datum_tijd'>,
<Axes: title={'right': "Stresses: ['RH_DE-BILT', 'EV24_DE-BILT']"}>,
<Axes: title={'center': 'Step response'}, xlabel='Time [days]'>,
<Axes: title={'left': 'Model Parameters ($n_c$=6)'}>]